Automated Data Extraction from Research Papers
PDF → Markdown → Metadata → Charts → Insight
Contents
Lecture, Practice, and Discussion for Week 6
- From PDF to Structured Data — Why and How
- Metadata extraction, PDF vs Markdown, and analysis strategies
- Build a PDF Data Extraction Pipeline (Streamlit)
- PDF → MD conversion, metadata charts, ontology, Q&A
- Week 5 Review & Midterm Progress Check
- From raw data to actionable research insights
The Data Problem in Research
You have PDFs. You need insights. The gap is enormous.
- Researchers accumulate hundreds of PDFs over a project
- Each PDF contains valuable data: authors, methods, findings, keywords
- But that data is trapped inside unstructured PDF format
- Manually extracting and organizing this data takes weeks
- PDF: designed for humans to read on screen/paper — visual layout, fonts, columns
- Database: designed for machines to query — structured, searchable, computable
- Getting from PDF to database requires extraction + structuring + validation
- This is exactly what LLMs are good at — understanding unstructured text
- Build a pipeline: PDF → Markdown (with metadata) → Charts → Q&A
- Week 5: you built a PDF viewer that chats about papers
- Week 6: you build a pipeline that extracts structured data from papers
- The difference: chat is ephemeral; extracted data is persistent and computable
PDF vs Markdown — Why Convert?
Understanding the fundamental format difference
| Aspect | Markdown | |
|---|---|---|
| Purpose | Visual presentation (print/screen) | Structured text (read/process) |
| Structure | Layout-based (coordinates, fonts) | Semantic (headings, lists, links) |
| AI-ready | Difficult (text extraction is lossy) | Easy (plain text with markup) |
| Searchable | Limited (no semantic structure) | Full text search + metadata |
| Editable | Requires special tools | Any text editor |
| Metadata | Embedded in binary format | YAML frontmatter (key-value) |
| Version control | Binary diff (useless) | Text diff (meaningful) |
| LLM-friendly | Must extract text first | Directly usable as context |
What Is Metadata?
Data about data — the key to unlocking your paper collection
- Metadata = structured information about a document (not the content itself)
- Title, authors, year, journal, keywords, DOI, methodology, findings
- Think of it as the index card for each paper in your collection
- With metadata from 100 papers, you can instantly answer:
- "How many papers use deep learning?" → keyword count
- "What's the publication trend over 5 years?" → year distribution
- "Who are the top authors in this field?" → author frequency
- "Which methods are most common?" → methodology count
- Without metadata, these questions require reading all 100 papers
Metadata in Markdown — YAML Frontmatter
Everything between --- markers is structured metadata, everything below is the document body
---
title: "Deep Learning for Material Property Prediction"
authors: ["Kim, J.", "Lee, S.", "Park, H."]
year: 2024
journal: "Nature Materials"
keywords: ["deep learning", "materials science", "property prediction"]
methodology: "Graph Neural Network"
key_findings: "GNN outperforms CNN by 15% on crystal property prediction"
---
## Abstract
This paper presents a novel approach to...LLM-Based Metadata Extraction
Using AI to read papers and extract structured data
- Feed raw PDF text to an LLM with a structured extraction prompt
- The LLM reads the paper and outputs JSON metadata + cleaned Markdown body
- This is a form of information extraction — one of AI's strongest capabilities
- You define what to extract — the LLM handles the how
- Default fields: title, authors, year, journal, keywords, abstract, methodology, findings
- Custom fields: you can add anything! ("sample_size", "equipment_used", "funding_source")
- The schema is your specification — same principle from Week 5
- LLM extraction is approximate — always verify critical metadata
- PDF text extraction is lossy — tables, figures, equations may be garbled
- Multi-column layouts and scanned PDFs are problematic
- Treat LLM-extracted metadata as draft requiring human validation (Week 2: hypothesis!)
(PyPDF2)"] B --> C["🧠 LLM
(extraction prompt)"] C --> D["📋 JSON Metadata"] C --> E["📝 Clean Markdown"] D --> F["💾 .md file
(YAML frontmatter)"] E --> F style C fill:#fff3e0,stroke:#f57c00 style F fill:#e8f5e9,stroke:#388e3c
The Full Extraction Pipeline
From folder of PDFs to structured database
(PyPDF2)"] B --> C["🧠 LLM +
schema"] C --> D["📋 Parse
JSON + body"] D --> E["💾 Save .md
(YAML frontmatter)"] E --> F["📁 MD Folder"] F --> G["📊 Charts"] F --> H["🕸️ Ontology"] F --> I["💬 Q&A"] style A fill:#fce4ec,stroke:#c62828 style C fill:#fff3e0,stroke:#f57c00 style F fill:#e8f5e9,stroke:#388e3c style G fill:#e1f5fe,stroke:#0288d1 style H fill:#f3e5f5,stroke:#7b1fa2 style I fill:#e1f5fe,stroke:#0288d1
What You Can Generate from Metadata
The real power — turning data into research intelligence
- Publication frequency by year → is the field growing or declining?
- Method evolution over time → which approaches are gaining traction?
- Trend prediction → based on current trajectory, what's next?
- Author frequency → who are the key researchers?
- Journal distribution → where is the field being published?
- Collaboration networks → who works with whom?
- Citation patterns → which papers are most referenced?
- Keyword frequency → what are the dominant topics?
- Keyword co-occurrence → which topics appear together?
- Topic correlation → how do subfields relate?
- Emerging keywords → new terms appearing in recent papers
- Field → Method → Finding hierarchy
- Research gap identification → what's NOT being studied?
- Cross-disciplinary connections → unexpected overlaps between fields
- Future direction synthesis → combining trends to predict opportunities
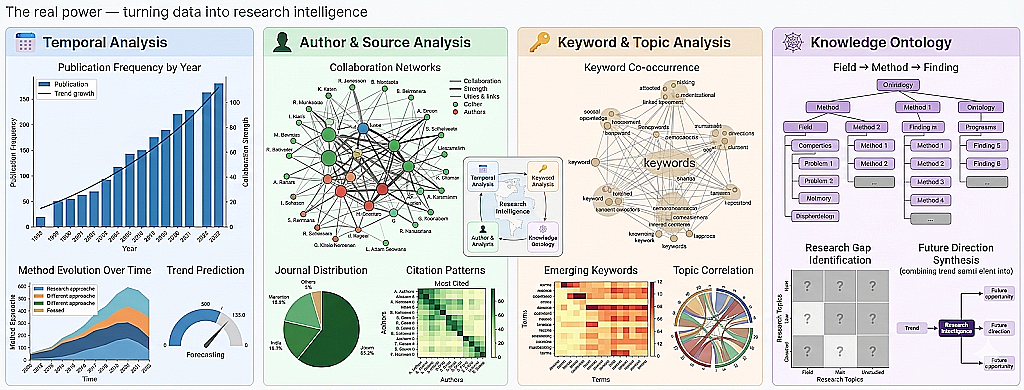
Example — What the Output Looks Like
These are generated automatically from extracted metadata
5 papers")) --- Lee(("🟠 Lee, S.
4 papers")) Kim --- Park(("🟡 Park, H.
3 papers")) Kim --- Chen(("🔵 Chen, W.
2 papers")) Lee --- Park Lee --- Tanaka(("🟢 Tanaka, Y.
3 papers")) Park --- Nguyen(("🟣 Nguyen, T.
2 papers")) Chen --- Tanaka style Kim fill:#ef4444,stroke:#b91c1c,color:#fff style Lee fill:#f97316,stroke:#c2410c,color:#fff style Park fill:#eab308,stroke:#a16207 style Tanaka fill:#22c55e,stroke:#15803d,color:#fff style Chen fill:#3b82f6,stroke:#1d4ed8,color:#fff style Nguyen fill:#a855f7,stroke:#7e22ce,color:#fff
From Data to Insight — Analysis Strategy
A 4-level framework for metadata-driven research intelligence
- Count, frequency, distribution of basic metadata fields
- "How many papers per year?", "What are the top 10 keywords?"
- Charts: bar charts, histograms, word clouds
- Tool: simple counting and grouping
- Cross-field comparisons, methodology vs outcome analysis
- "Do DL papers cite more than traditional ML papers?"
- "How do methods differ between journals?"
- Tool: pivot tables, grouped bar charts
- Co-occurrence networks, citation graphs, topic correlation
- "Which keywords always appear together?"
- "Which authors bridge two research communities?"
- Tool: network graphs, ontology diagrams
- Trend extrapolation, gap identification, opportunity mapping
- "Based on 5 years of data, what will the hot topics be in 2027?"
- "Where are the unexplored intersections between fields?"
- Tool: LLM-powered analysis + human judgment
The Hidden Challenge — Data Normalization
LLMs extract text, but the same entity can appear in many forms
- "Kim, J." / "J. Kim" / "Joon Kim" / "KIM, JOON" → same person!
- "Lee, S." — is it Sungmin Lee or Soyoung Lee?
- Without normalization, your author network has duplicate nodes
- Same issue for: journal names, keywords, institutions
- Case normalization: "KIM, SOYEON" → "Soyeon Kim"
- Name order: "Kim, Soyeon" → "Soyeon Kim" (unify to First Last)
- Punctuation cleanup: remove dots, hyphens→space, trim brackets
- Keyword unification: "deep learning" / "Deep Learning" / "DL" → "deep learning"
normalize_author()- Input: any author name format from the LLM
- Output: consistent full name in Title Case
- Applied before counting, charting, and network building
- Same pattern works for keywords, journals, etc.
Customizing the Extraction Schema
The schema defines what data you get — tailor it to YOUR research
# Default schema (works for most papers)
- title: Paper title
- authors: List of author names
- year: Publication year
- journal: Journal or conference name
- keywords: Key phrases
- abstract: The abstract
- methodology: Primary research method
- key_findings: 1-2 sentence summary
# Example: Materials Science custom fields
- material_system: Primary material studied
- synthesis_method: How the material was made
- characterization_tools: Equipment used (XRD, SEM, TEM, etc.)
- performance_metric: Key performance number and unit
# Example: Machine Learning custom fields
- dataset: Dataset used for training/evaluation
- model_architecture: Neural network architecture
- baseline_comparison: What was compared against
- accuracy_metric: Best reported accuracy/F1/BLEU score- Think: what metadata would make YOUR literature review easier?
- Add fields specific to your field — the LLM will try to extract them
- More specific schema → more useful metadata → better analysis
- This is prompt engineering (Week 3) applied to data extraction
Lecture Summary — From PDF to Insight
Key takeaways
- PDFs trap data in visual format; Markdown makes it AI-ready
- YAML frontmatter stores structured metadata alongside the content
- LLM extracts metadata from raw text — approximate but powerful
- With structured metadata, you can count, compare, connect, and predict
- 4 levels: Descriptive → Comparative → Relational → Predictive
- Levels 1-3 are computation; Level 4 is human judgment
- Week 5: you built a PDF viewer (read and chat)
- Week 6: you build a PDF pipeline (extract, structure, analyze)
- The difference: persistent, queryable, computable data
Part 2: Practice
Build a PDF Data Extraction Pipeline — Streamlit Web App
What We'll Build Today
A 4-tab Streamlit app for end-to-end paper analysis
- Tab 1 — Extract: Upload PDFs → LLM extracts metadata → saves as
.mdfiles - Tab 2 — Metadata Table: View all extracted metadata as a table + download CSV
- Tab 3 — Charts & Ontology: Year/field/method distributions + keyword network
- Tab 4 — Q&A: Chat about your paper collection using extracted data
- Streamlit — web framework (same as Week 5)
- PyPDF2 — PDF text extraction
- Pandas — data tables and CSV export
- OpenAI client — LLM calls (Gemini / Ollama)
app.py— Main Streamlit app (4 tabs)pdf_to_md.py— PDF → Markdown converter with LLM extractionllm_client.py— LLM client (extraction + chat)chart_generator.py— Charts and ontology from metadatapdfs/— input folder (put your PDFs here)md_output/— output folder (generated .md files)
Step 0 — Setup
Install dependencies and prepare folders
cd practices/week_06
pip install streamlit PyPDF2 openai python-dotenv pandaspractices/week_06/
app.py # Main Streamlit app
pdf_to_md.py # PDF → MD conversion
llm_client.py # LLM client
chart_generator.py # Visualization generators
pdfs/ # Put your PDF files here
md_output/ # Generated .md files (auto-created)- Copy
.envfrom Week 5 (orpractices/.env) - Same Gemini / Ollama setup — no new configuration needed
- Prepare 3-5 PDF papers from your research for testing
Step 1 — PDF → Markdown Converter (pdf_to_md.py)
Extract text, send to LLM, parse response, save as .md
# pdf_to_md.py (key functions)
from PyPDF2 import PdfReader
import json, os
def extract_pdf_text(pdf_path):
reader = PdfReader(pdf_path)
return "\n".join(p.extract_text() or "" for p in reader.pages)
def build_extraction_prompt(raw_text, user_schema):
return f"""Extract metadata as JSON + body as Markdown.
## Schema: {user_schema}
## Output: ```json {{ ... }} ``` ---BODY--- (markdown body)
## Paper Text: {raw_text[:30000]}"""
def parse_llm_response(response_text):
metadata = {}
if "```json" in response_text:
json_str = response_text.split("```json")[1].split("```")[0]
metadata = json.loads(json_str)
body = response_text.split("---BODY---")[1] if "---BODY---" in response_text else ""
return metadata, body.strip()
def save_markdown(output_dir, filename, metadata, body):
"""Save as .md with YAML frontmatter (lists as indented items)."""
lines = ["---"]
for k, v in metadata.items():
if isinstance(v, list):
lines.append(f"{k}:")
for item in v: lines.append(f' - "{item}"')
else:
lines.append(f'{k}: "{v}"')
lines.extend(["---", "", body])
md_path = os.path.join(output_dir, filename.replace(".pdf", ".md"))
with open(md_path, "w", encoding="utf-8") as f:
f.write("\n".join(lines))
return md_path
def load_all_metadata(md_dir):
"""Load YAML frontmatter from all .md files (supports list fields)."""
all_meta = []
for fname in sorted(os.listdir(md_dir)):
if not fname.endswith(".md"): continue
with open(os.path.join(md_dir, fname), encoding="utf-8") as f:
content = f.read()
if not content.startswith("---"): continue
parts = content.split("---", 2)
meta, current_key, current_list = {"_filename": fname}, None, []
for line in parts[1].strip().split("\n"):
if line.startswith(" - "): # list item
current_list.append(line.strip()[2:].strip('"'))
else:
if current_key and current_list: # save previous list
meta[current_key] = json.dumps(current_list)
current_list = []
if ": " in line:
k, v = line.split(": ", 1)
v = v.strip().strip('"')
if v in ("", "|"): current_key = k
else: meta[k] = v; current_key = None
elif line.endswith(":"): # key with list below
current_key = line[:-1].strip()
else: current_key = None
if current_key and current_list:
meta[current_key] = json.dumps(current_list)
all_meta.append(meta)
return all_metaStep 2 — LLM Client (llm_client.py)
Two functions — one for extraction, one for Q&A chat
# llm_client.py
from openai import OpenAI
import os
from dotenv import load_dotenv
load_dotenv()
def get_client(provider="Gemini"):
# Same as Week 5 — Gemini / Ollama / OpenAI
...
def extract_metadata(client, model, raw_text, prompt):
"""One-shot extraction — no streaming needed."""
response = client.chat.completions.create(
model=model,
messages=[
{"role": "system", "content": "You are a precise metadata extractor."},
{"role": "user", "content": prompt}
],
max_tokens=4096,
)
return response.choices[0].message.content
def chat_with_data(client, model, context, user_message, history):
"""Chat about extracted data — streaming."""
system_msg = f"""You are a research data analyst.
Use the paper metadata below to answer questions.
# Paper Data\n{context}"""
messages = [{"role": "system", "content": system_msg}]
messages.extend(history)
messages.append({"role": "user", "content": user_message})
return client.chat.completions.create(
model=model, messages=messages, stream=True)- Extraction (Tab 1): single call, no streaming, structured output →
extract_metadata() - Q&A (Tab 4): multi-turn chat, streaming, conversational →
chat_with_data() - Same LLM, different system prompts → different behavior (Week 3 principle!)
Step 3 — Chart Generator (chart_generator.py)
Turn metadata into visualizations
# chart_generator.py
from collections import Counter
import json, re
def normalize_author(name):
"""Normalize author name — keep full name, clean formatting."""
name = name.strip().strip('"').replace(".", "").replace("-", " ")
name = re.sub(r"[\d()\[\]{}*]", "", name)
name = " ".join(name.split())
if "," in name: # "Last, First" → "First Last"
parts = [p.strip() for p in name.split(",", 1)]
name = f"{parts[1]} {parts[0]}".strip()
return name.title()
def normalize_authors_list(authors_raw):
"""Parse and normalize author list from metadata."""
if isinstance(authors_raw, list): names = authors_raw
elif isinstance(authors_raw, str):
try: names = json.loads(authors_raw.replace("'", '"'))
except: names = [n.strip() for n in authors_raw.split(",")]
else: return []
return [normalize_author(n) for n in names if n.strip()]
def author_cooccurrence(metadata_list):
"""Build co-authorship edges + author counts."""
edges, author_count = Counter(), Counter()
for meta in metadata_list:
authors = normalize_authors_list(meta.get("authors", "[]"))
for a in authors: author_count[a] += 1
for i, a1 in enumerate(authors):
for a2 in authors[i+1:]:
edges[tuple(sorted([a1, a2]))] += 1
return {"edges": [{"source": s, "target": t, "weight": w}
for (s, t), w in edges.most_common(30)],
"counts": dict(author_count.most_common(20))}
def count_by_field(metadata_list, field):
"""Count occurrences of a metadata field."""
counter = Counter()
for meta in metadata_list:
counter[meta.get(field, "Unknown")] += 1
return dict(counter.most_common(20))
def year_distribution(metadata_list):
"""Count papers per year."""
counter = Counter()
for meta in metadata_list:
try: counter[int(meta.get("year", 0))] += 1
except: pass
return dict(sorted(counter.items()))
def build_keyword_cooccurrence(metadata_list):
"""Build keyword co-occurrence pairs."""
# ... (same pattern as author_cooccurrence)
def generate_mermaid_ontology(metadata_list, center_topic="Research"):
"""Generate Mermaid graph from fields, methods, keywords."""
# ... graph TD with CENTER → top fields/methods/keywords
def generate_mermaid_author_network(metadata_list):
"""Generate Mermaid graph showing author collaboration network."""
data = author_cooccurrence(metadata_list)
lines = ["graph LR"]
for i, (author, cnt) in enumerate(data["counts"].items()):
lines.append(f' A{i}(("{author}<br>{cnt} papers"))')
for edge in data["edges"]:
# Connect co-authors with --- edges
...
return "\n".join(lines)Step 4 — Tab 1: Extract (app.py)
Upload PDFs → LLM extracts metadata → saves as .md files
# app.py — Tab 1 (Extract)
with tab1:
st.header("PDF → Markdown Extraction")
pdf_files = os.listdir(PDF_DIR) # List PDFs in folder
md_files = os.listdir(MD_DIR) # List already-extracted MDs
# Show status per PDF
for pdf_name in pdf_files:
md_name = pdf_name.replace(".pdf", ".md")
status = "✅" if md_name in md_files else "⏳"
st.markdown(f"- `{pdf_name}` — {status}")
if st.button("🚀 Extract All Pending"):
progress = st.progress(0)
for i, pdf_name in enumerate(pending_pdfs):
progress.progress(i / len(pending_pdfs), f"Extracting: {pdf_name}")
raw_text = extract_pdf_text(os.path.join(PDF_DIR, pdf_name))
prompt = build_extraction_prompt(raw_text, user_schema)
response = extract_metadata(client, model, raw_text, prompt)
metadata, body = parse_llm_response(response)
save_markdown(MD_DIR, pdf_name, metadata, body)
st.rerun()- Unlike Week 5 (one PDF at a time in chat), Week 6 processes all PDFs in a pipeline
- Progress bar shows extraction status for each file
- Results persist in
md_output/— no re-extraction needed on reload
Step 5 — Tab 2: Table & Tab 3: Charts
View, export, and visualize extracted metadata
# app.py — Tab 2 (Metadata Table)
with tab2:
all_meta = load_all_metadata(MD_DIR)
df = pd.DataFrame(all_meta)
st.dataframe(df, use_container_width=True)
st.download_button("📥 Download CSV", df.to_csv(), "metadata.csv")
# app.py — Tab 3 (Charts & Ontology) — uses altair for horizontal bar charts
import altair as alt
with tab3:
# Year distribution (horizontal bar chart)
year_data = year_distribution(all_meta)
df = pd.DataFrame(year_data.items(), columns=["Year", "Count"])
df["Year"] = df["Year"].astype(str)
chart = alt.Chart(df).mark_bar().encode(
y=alt.Y("Year:N", sort="-x"), x="Count:Q")
st.altair_chart(chart, use_container_width=True)
# Author collaboration network (normalized full names)
author_mermaid = generate_mermaid_author_network(all_meta)
st.code(author_mermaid, language="mermaid")
author_data = author_cooccurrence(all_meta)
st.dataframe(pd.DataFrame(author_data["counts"].items(),
columns=["Author", "Papers"]))
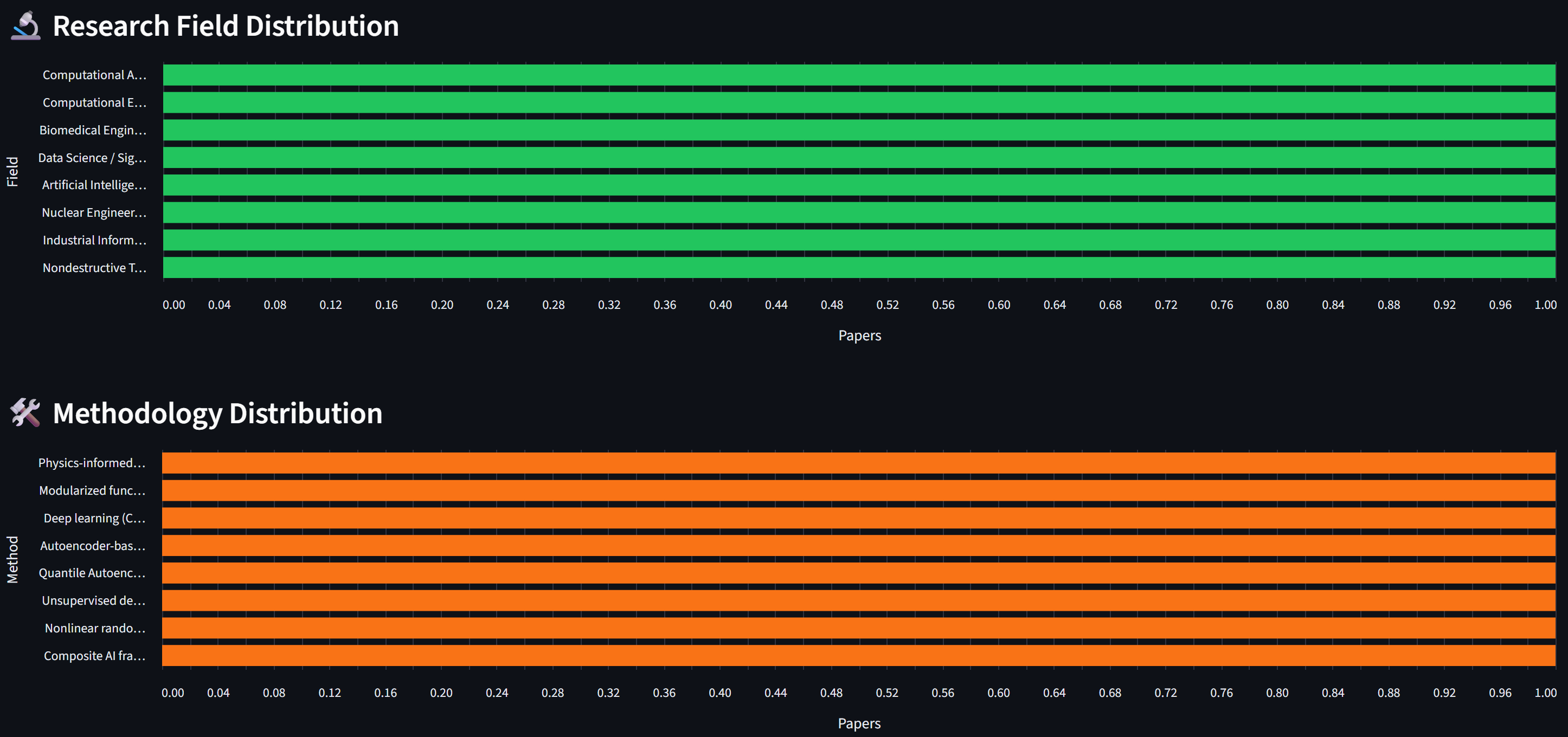
# Field / Methodology distribution (horizontal bar charts)
for field, label in [("research_field", "Field"), ("methodology", "Method")]:
data = count_by_field(all_meta, field)
df = pd.DataFrame(data.items(), columns=[label, "Count"])
chart = alt.Chart(df).mark_bar().encode(
y=alt.Y(f"{label}:N", sort="-x"), x="Count:Q")
st.altair_chart(chart, use_container_width=True)
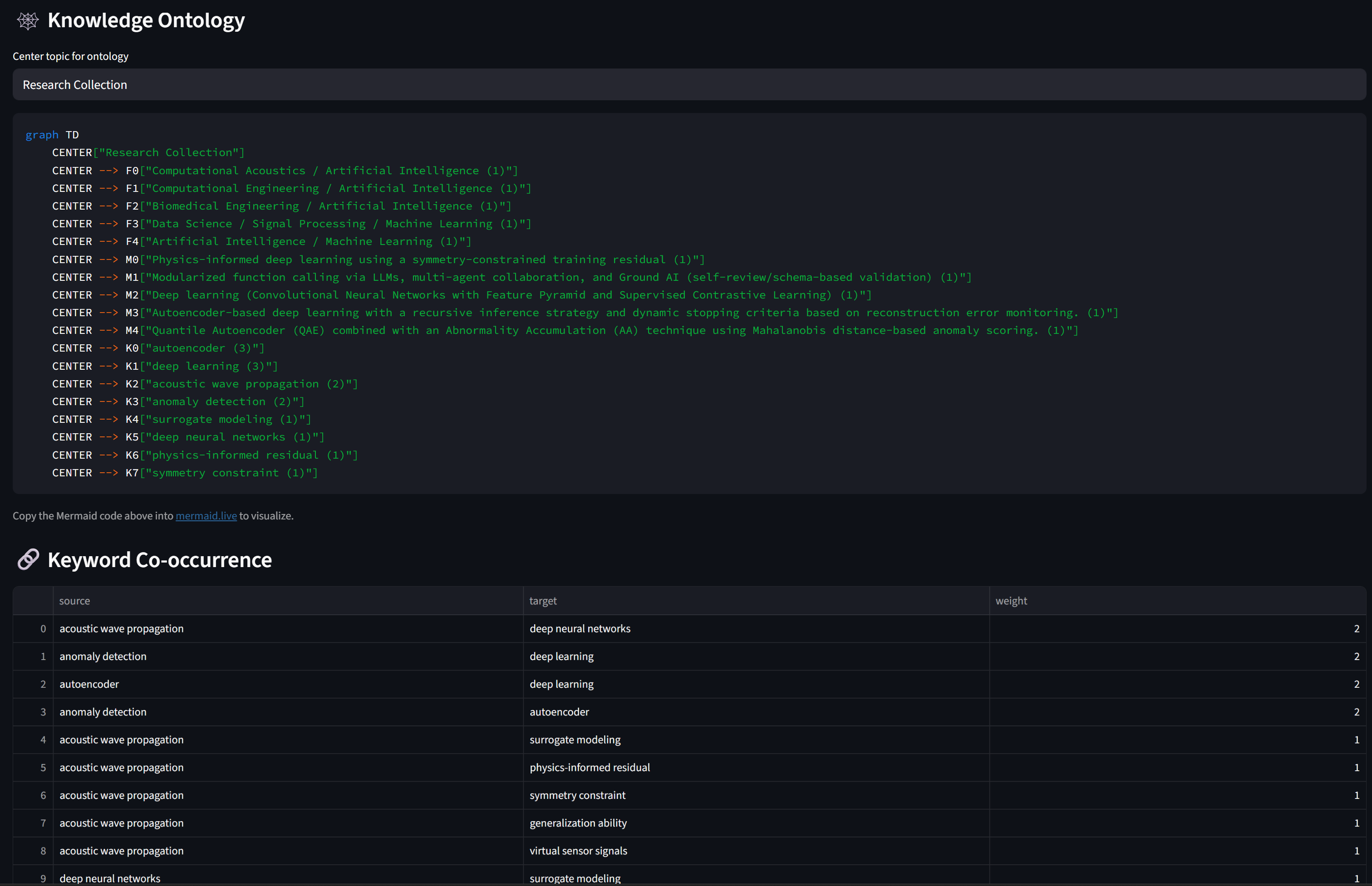
# Knowledge ontology + Keyword co-occurrence (same as before)
st.code(generate_mermaid_ontology(all_meta), language="mermaid")
st.dataframe(pd.DataFrame(build_keyword_cooccurrence(all_meta)))Step 6 — Tab 4: Q&A on Your Paper Collection
Chat with AI about all your extracted papers
# app.py — Tab 4 (Q&A)
with tab4:
# Build context from all metadata
context = "\n\n".join(
f"### {m.get('title')}\n" +
"\n".join(f"- **{k}**: {v}" for k, v in m.items())
for m in all_meta
)
# Quick analysis buttons
quick_prompts = {
"📊 Trend Analysis": "Analyze temporal trends...",
"🔍 Common Themes": "What common themes...",
"⚡ Research Gaps": "Identify 3-5 gaps...",
"🔀 Cross-Pollination": "Find unexpected connections...",
}
for label, template in quick_prompts.items():
if st.button(label):
# Send template as prompt → stream response
...
# Chat input (same streaming pattern as Week 5)
if prompt := st.chat_input("Ask about your papers..."):
stream = chat_with_data(client, model, context, prompt, history)
for chunk in stream:
... # stream to placeholder- Week 5: injected full PDF text into system prompt (huge, limited to 2-3 papers)
- Week 6: injects extracted metadata (compact, scales to 50+ papers)
- Structured metadata = more papers in less tokens = better analysis
Full Pipeline — Architecture
From raw PDFs to interactive research intelligence
Step 7 — Run and Test
Launch the app and process your papers
cd practices/week_06
streamlit run app.pyExpected UI (4 tabs):
┌──────────────────────────────────────────────────────┐
│ [1️⃣ Extract] [2️⃣ Metadata Table] [3️⃣ Charts] [4️⃣ Q&A] │
│ │
│ Tab 1 — PDF → Markdown Extraction │
│ │
│ 📁 PDF Files: │
│ - paper1.pdf — ✅ Extracted │
│ - paper2.pdf — ✅ Extracted │
│ - paper3.pdf — ⏳ Pending │
│ │
│ [🚀 Extract All Pending (1)] [🔄 Re-extract All] │
│ │
│ Processing Log: │
│ ✅ paper1.pdf → paper1.md │
│ ✅ paper2.pdf → paper2.md │
└──────────────────────────────────────────────────────┘- Start with 3-5 short papers (< 10 pages each) for faster extraction
- Check
md_output/folder to verify YAML frontmatter is correct - If metadata looks wrong, try editing the schema in the sidebar for clearer instructions
- Gemini works best for extraction (large context window)
Screen Shots of the App
Views of the app and process your papers
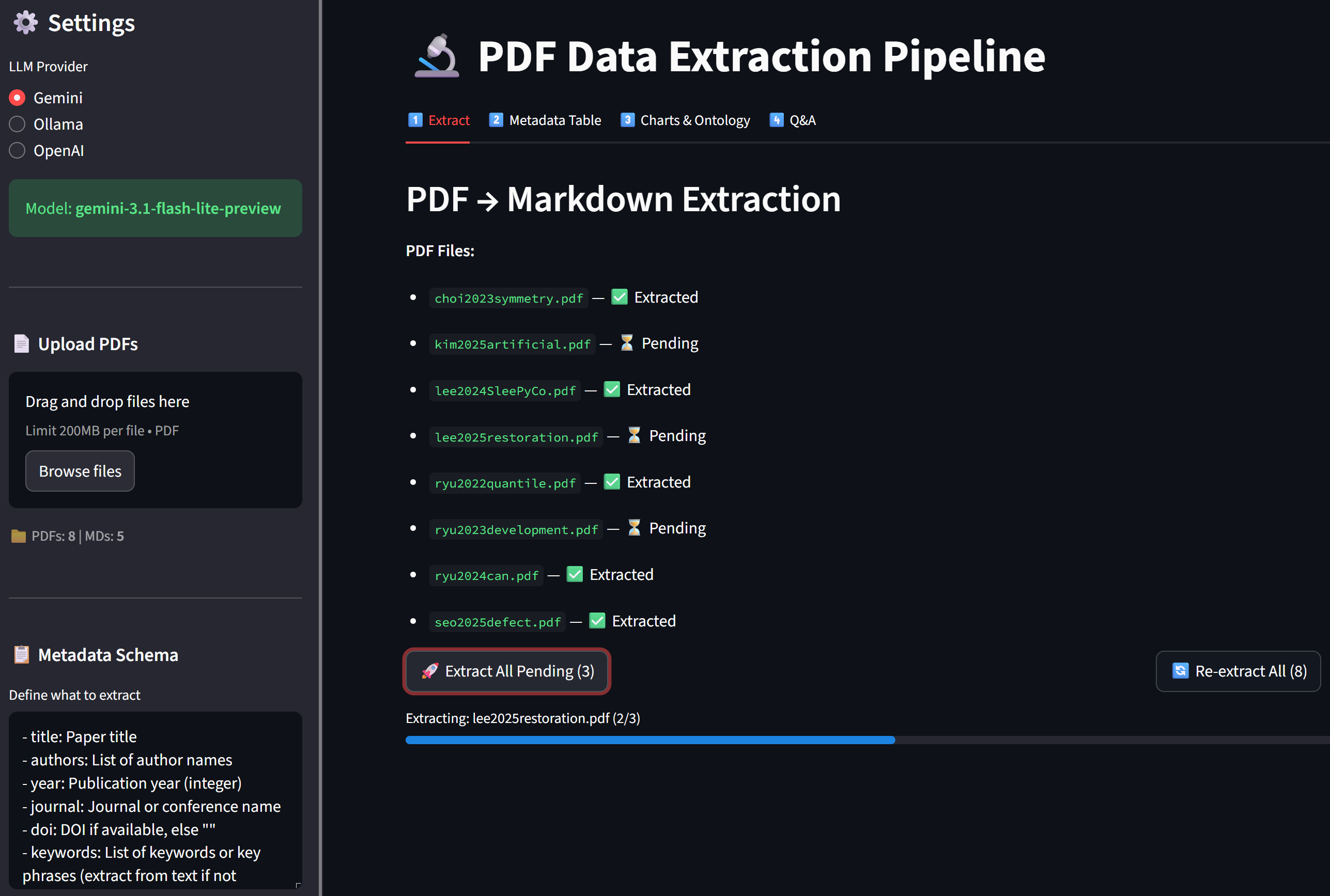

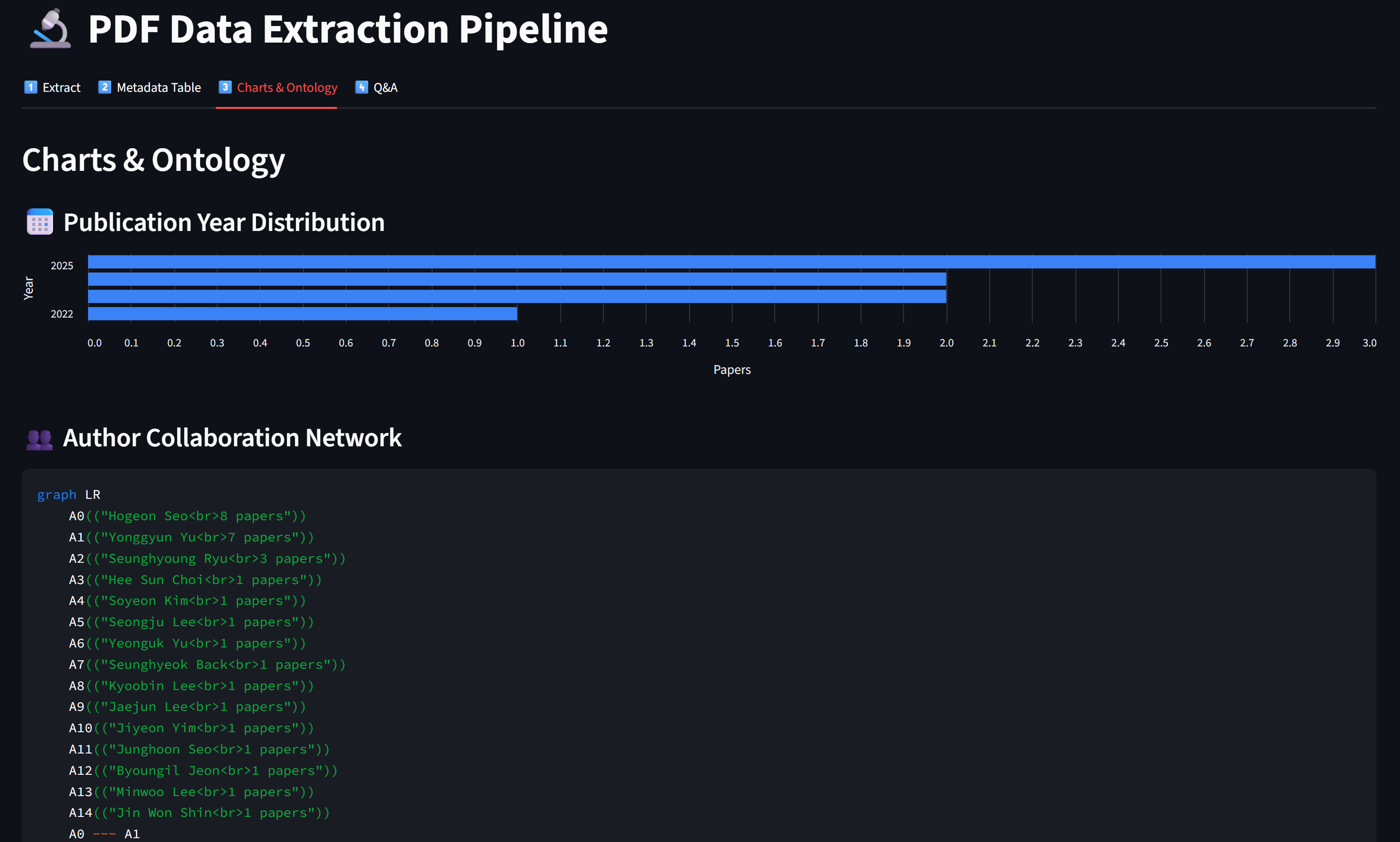
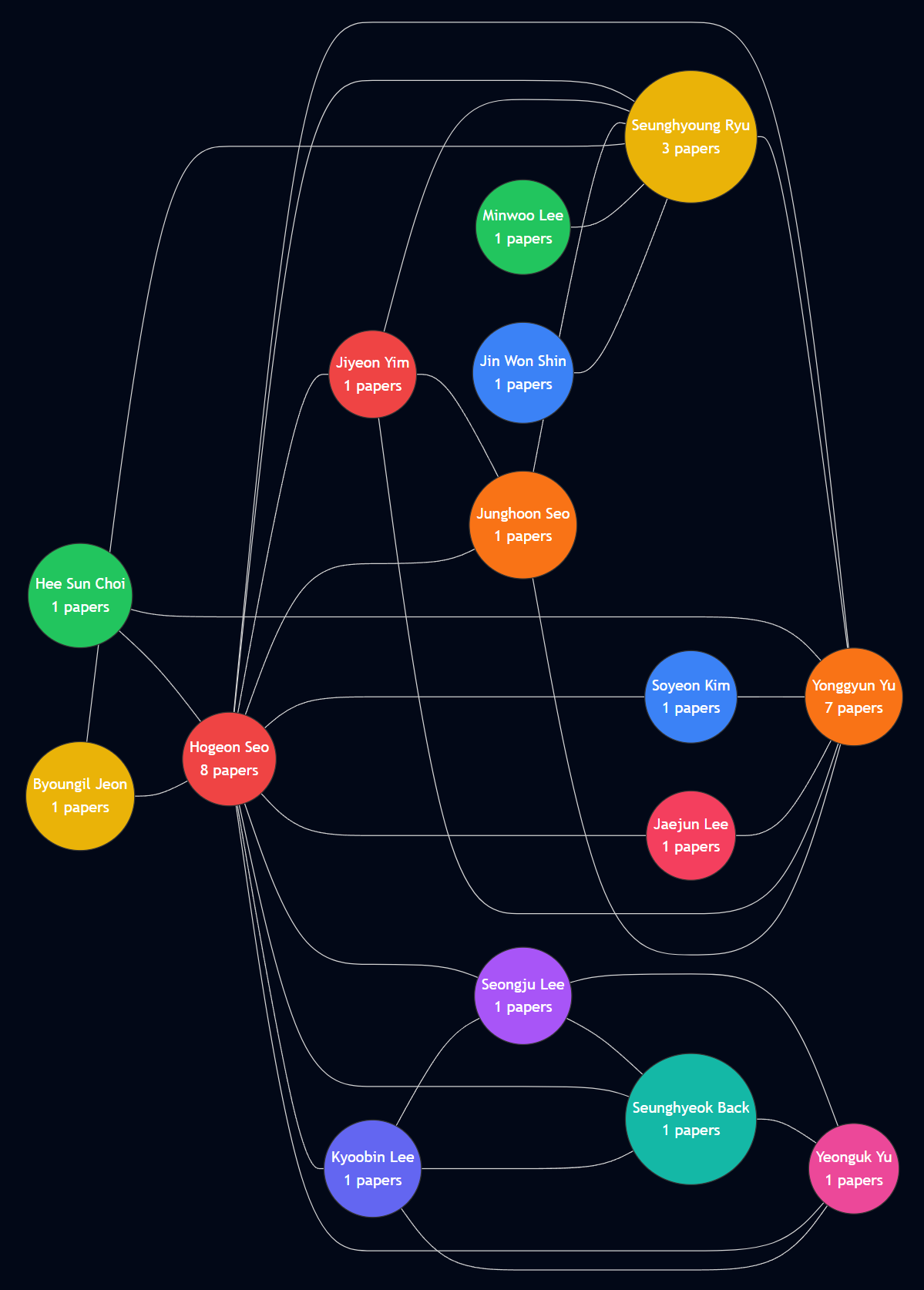
✅ Practice Checklist
Complete these tasks during the hands-on session
- [ ] Set up
.envand prepare 3-5 PDF papers inpdfs/folder - [ ] Create all 4 files:
app.py,pdf_to_md.py,llm_client.py,chart_generator.py - [ ] Run
streamlit run app.pyand verify the 4-tab UI loads - [ ] Tab 1: Extract all PDFs → verify
.mdfiles appear inmd_output/ - [ ] Open a
.mdfile and verify YAML frontmatter has correct metadata - [ ] Tab 2: View metadata table → download CSV
- [ ] Tab 3: Check year/field/method charts → view ontology Mermaid code
- [ ] Tab 4: Ask a question about your paper collection → verify AI references specific papers
- [ ] (Bonus) Customize the schema for your research field and re-extract
- [ ] (Bonus) Use a quick analysis button (Trend Analysis, Research Gaps)
- [ ] (Bonus) Compare results between Gemini and Ollama
Part 3: Discussion
Core Competency & Midterm Progress
Core Competency — The "Irreplaceable" Researcher
"In an era where AI conducts experiments and writes papers, what is the irreplaceable skill of a researcher?"
- "The irreplaceable core competency is acting as the visionary architect of the blueprint"
- "Let the algorithms grind through the data swamps; human genius is strictly reserved for asking the universe-breaking questions"
- Key idea: problem selection > problem solving
- "The irreplaceable core competency is the unwavering moral integrity and personal accountability required to seek the objective truth"
- "Relying on AI as a crutch threatens to erode the fundamental critical thinking and honest hard work"
- Key idea: ethical responsibility > speed
- "The irreplaceable core competency is rigorous, skeptical validation — acting as the ultimate fail-safe against unchecked automated systems"
- "We must mandate exhaustive human oversight at every procedural juncture"
- Key idea: verification > trust
Three Views, One Question — Where Do You Stand?
Each AI agent reflects a real debate in the research community
- Iron Man says: aim the blast — your value is in choosing what to research
- Captain America says: hold the line — your value is in doing it honestly
- Hulk says: check the output — your value is in catching AI's mistakes
- Are these complementary or contradictory?
- Which view best matches your daily research reality?
- Has AI already changed what skills matter in YOUR field?
- What skill do you use that you believe AI cannot learn?
Your Responses — The Class Map
17 responses — most of you agreed with all three, but the interesting part is WHERE you disagreed
- All three (1+2+3): Huy, Yadanar, Irfan, Waad, Tan, DongYun, Nazhiefah — "all three are complementary"
- Iron Man + Hulk (1+3): Seher, Gyeongsu, Minh — "vision + validation, skip the lecture on virtue"
- Captain America + Hulk (2+3): Manuella, Ly — "judgment + ethics, not just big ideas"
- Hulk only (3): Namcheol, Rupam, Hyunwoo — "validation is THE core skill"
- Captain America (2): Margareth — "ethics is hardest to replace"
- Han: "All three are partially right and collectively miss the point"
- Introduced a new concept: "cultivated epistemic taste"
- "That taste only develops through doing the work AI is now replacing"
- This challenges the entire premise — are we losing the very skill we claim is irreplaceable?
Theme 1 — Hulk Wins the Vote
Skeptical validation emerged as the class's top priority
- Every single response included validation as essential — Hulk's view was universally endorsed
- Jaewhoon: "Computer is deterministic, AI is probabilistic — that's why we must check"
- Namcheol: "The researcher must know when to pull the emergency stop button"
- Hyunwoo: "In robotics, we must cross-reference AI outputs against physical reality"
- Rupam: "Skepticism is intrinsic — it's not something that can be easily taught or programmed"
- Minh: "Strategic Synthesis — bridging 'what can be simulated' and 'what should be built'"
- Huy: "We do not need to compete with AI but rather judge it"
- The class is saying: validation isn't just checking boxes — it requires deep domain knowledge
Theme 2 — The Captain America Debate
The most divisive AI agent — is "doing things the hard way" valuable or outdated?
- Gyeongsu: "Efficiency isn't a moral flaw. Sticking to manual labor just for tradition is a speed bump"
- Rupam: "We cannot dismiss AI because it is fast — that argument feels illogical"
- Both argue: integrity matters, but Captain America wrongly equates slowness with virtue
- Margareth: "Ethics and moral integrity are the aspects hardest to replace with AI"
- Nazhiefah: "Rather than speed everything, the process of research itself is the important thing"
- Ly: "The researcher must act as an ethical anchor and guardian of scientific validity"
- Han resolved the tension: "epistemic taste only develops through doing the work AI is replacing — struggling through calculations, debugging by hand, reading papers slowly"
- This reframes Captain America: it's not about virtue — it's about building the judgment you need
- If you skip the hard work, you lose the ability to validate (connecting Theme 1 and 2)
Theme 3 — Han's Challenge
"If you outsource the friction, you become someone who can't read the blueprints"
- Han's concept: the judgment to know which questions are worth asking, which results matter
- "It's built from years of wrestling directly with a domain's hardest problems"
- "AI can optimize within a search space; it cannot reliably define the right search space"
- The skill we need most (judgment) is built through the work we're outsourcing to AI
- Nazhiefah echoed this: "we sometimes want quick results but are lazy to do boring stuff"
- Jaewhoon: like a grad assistant that hallucinates — if the PI doesn't check, the PI takes the blame
- Question: how do we develop epistemic taste if AI does all the "boring" foundational work?
🗣️ Reflect — Apply Han's Paradox to Your Midterm
5 minutes — Think about this tension in YOUR app
- Your midterm app automates some research task with AI
- Does your app preserve the user's ability to develop judgment?
- Or does it outsource the "friction" that builds expertise?
- Is there a way to design for both efficiency AND skill development?
- Consider adding a "learning mode" vs "production mode" to your app
- Learning mode: shows the AI's reasoning, asks user to verify steps
- Production mode: runs autonomously for experienced users
- This resolves Han's paradox — the tool helps you learn AND speeds you up
Midterm Progress Check
Where should you be right now?
- Decided on your project topic and problem statement
- Drafted the 5-question specification document
- Identified which LLM provider and framework you'll use
- Refine your spec based on feedback
- Start building your prototype — even a skeleton UI is progress
- Consider: can you use today's extraction pipeline pattern?
- Week 7: Specification document due (submit on LMS)
- Week 8: Working prototype + 5-minute live demo
- Submit by April 17 (Fri) 24:00 → email to hogeony@ust.ac.kr
- Two weeks left — start coding NOW if you haven't already
🗣️ Week 6 Discussion Questions (UST LMS)
Post your response on the forum this week
1. Core Competency: Read the three AI agent opinions above. Which perspective do you most agree with, and why? What would you add that none of the three agents mentioned? Think about your own research field — what specific skill makes you irreplaceable?
2. Today you learned to extract structured metadata from PDFs using LLM. Design a custom extraction schema for your specific research field: what 5-8 metadata fields would be most valuable for analyzing papers in YOUR domain? Why these fields?
3. Midterm progress update: Share your current specification status. What's your app's name, core problem, and 3 main features? What's your biggest design challenge so far?
Want to Learn More?
Data Extraction & Processing
📚 PyPDF2 Documentation 📚 Marker — Best PDF to Markdown Converter 📚 LangChain Document Loaders
Metadata & Knowledge Graphs
📚 Semantic Scholar API 📚 OpenAlex — Open Research Knowledge Graph 📚 YAML Frontmatter Specification
Visualization
📚 Mermaid.js Documentation 📚 Streamlit Charts API 📚 Plotly for Python
Anthropic Free Online Courses (Recommended)
Wrap-Up of Week 6
Three things to remember
- PDF traps data in visual format; Markdown + YAML frontmatter makes it AI-ready; LLM extracts metadata from raw text; 4-level analysis framework (describe → compare → connect → predict)
- Built a 4-tab extraction pipeline: PDF→MD extraction, metadata table, charts/ontology, and Q&A; metadata scales to 50+ papers where full-text chat cannot
- Core Competency: Hulk (validation) won the vote; Captain America was most divisive; Han's "epistemic taste" paradox — outsourcing friction destroys judgment; midterm due April 17 24:00 via email
Next week: Specification document due — finalize your project design and prepare for prototyping in Week 8.